Factorial Design
Overview
Factorial designs are experimental designs used to study the effects of two or more factors simultaneously. They allow estimation of main effects and interactions between factors, making them efficient and informative.
When to Use
- Studying multiple factors at the same time
- Detecting interactions between factors
- Efficient use of experimental units
Setting Up Factorial Design with speed
Now we can create a data frame representing a factorial design. Note that the treatment column we are creating is the interaction (or combination) of the individual treatments.
treatment_a <- paste0("A", 1:8)
treatment_b <- paste0("B", 1:3)
treatments <- with(expand.grid(treatment_a, treatment_b), paste(Var1, Var2, sep = "-"))
factorial_design <- initialise_design_df(treatments, 24, 3, 8, 3)
head(factorial_design) row col treatment row_block col_block block
1 1 1 A1-B1 1 1 1
2 2 1 A2-B1 1 1 1
3 3 1 A3-B1 1 1 1
4 4 1 A4-B1 1 1 1
5 5 1 A5-B1 1 1 1
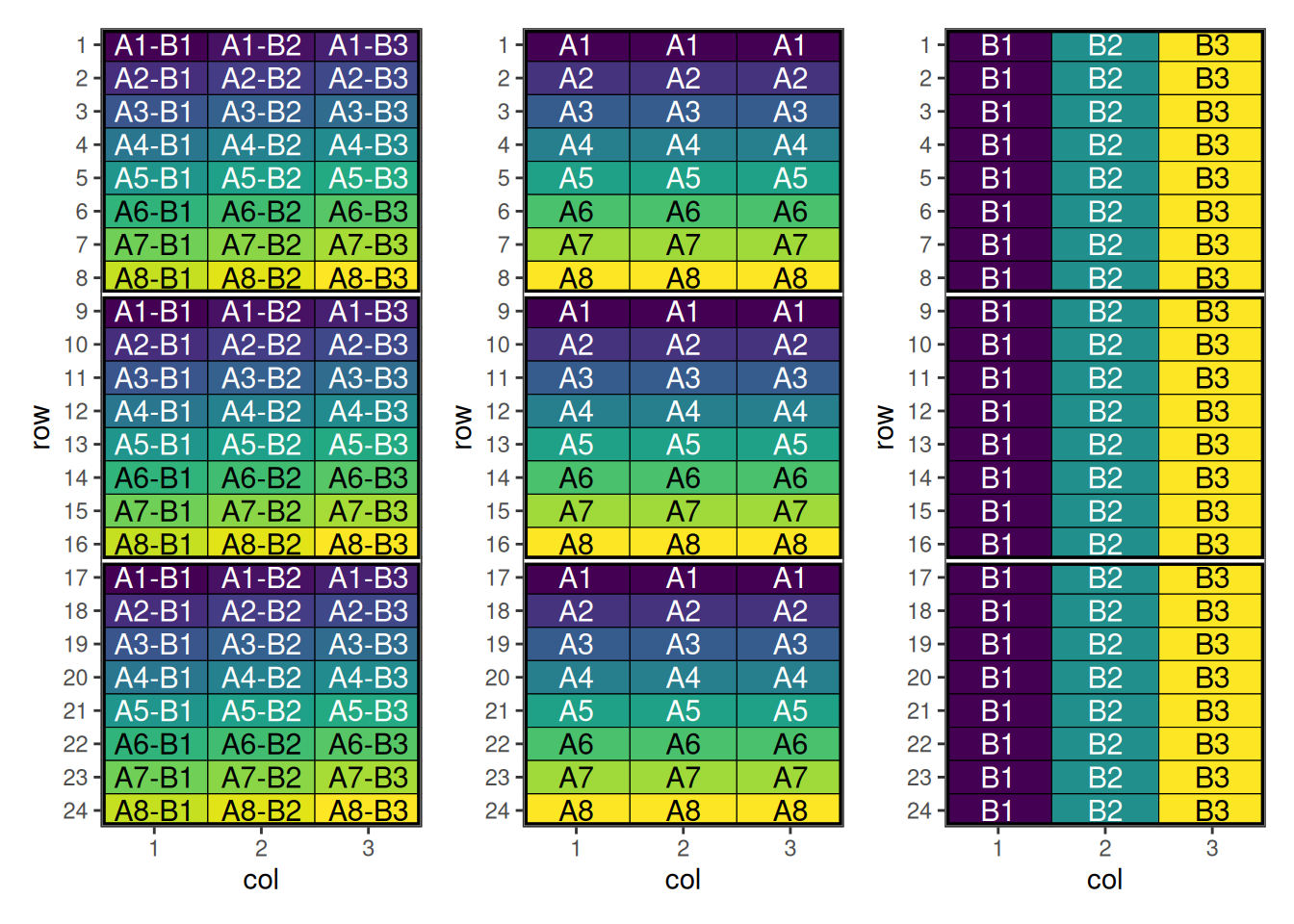
6 6 1 A6-B1 1 1 1Plotting these factors shows the initial layout of the interaction and main effects.
Performing the Optimisation
For factorial designs, speed provides a customised objective function objective_function_factorial which allows us to pass interaction_weight and main_weight arguments to control the spatial balance of those effects. The optimise_params argument is also used in this case to adjust the optimisation strategy due to the difficulty of optimising such designs.
Make sure factorial_separator matches how you constructed the interaction treatment (here we used "-"). If your treatments use a different separator (e.g. "A1:B2"), pass factorial_separator = ":".
optimise_params <- optim_params(
swap_count = 3,
random_initialisation = 10,
adaptive_swaps = TRUE,
swap_all_blocks = TRUE,
cooling_rate = 0.999
)
factorial_result <- speed(
data = factorial_design,
swap = "treatment",
swap_within = "block",
spatial_factors = ~ row + col,
obj_function = objective_function_factorial,
optimise_params = optimise_params,
early_stop_iterations = 10000,
iterations = 200000,
interaction_weight = 10,
seed = 112
)row and col are used as row and column, respectively.Optimising level: single treatment within block
Level: single treatment within block Iteration: 1000 Score: 158.118 Best: 105.3975 Since Improvement: 174
Level: single treatment within block Iteration: 2000 Score: 123.3043 Best: 105.3975 Since Improvement: 1174
Level: single treatment within block Iteration: 3000 Score: 121.795 Best: 100.6957 Since Improvement: 261
Level: single treatment within block Iteration: 4000 Score: 104.8199 Best: 93.92547 Since Improvement: 202
Level: single treatment within block Iteration: 5000 Score: 65.14907 Best: 58.38509 Since Improvement: 140
Level: single treatment within block Iteration: 6000 Score: 51.15528 Best: 51.15528 Since Improvement: 89
Level: single treatment within block Iteration: 7000 Score: 42.10559 Best: 42.10559 Since Improvement: 381
Level: single treatment within block Iteration: 8000 Score: 40.81988 Best: 40.81988 Since Improvement: 501
Level: single treatment within block Iteration: 9000 Score: 39.81988 Best: 39.81988 Since Improvement: 742
Level: single treatment within block Iteration: 10000 Score: 36.81988 Best: 36.81988 Since Improvement: 537
Level: single treatment within block Iteration: 11000 Score: 36.81988 Best: 36.81988 Since Improvement: 1537
Level: single treatment within block Iteration: 12000 Score: 36.81988 Best: 36.81988 Since Improvement: 2537
Level: single treatment within block Iteration: 13000 Score: 36.81988 Best: 36.81988 Since Improvement: 3537
Level: single treatment within block Iteration: 14000 Score: 36.81988 Best: 36.81988 Since Improvement: 4537
Level: single treatment within block Iteration: 15000 Score: 36.81988 Best: 36.81988 Since Improvement: 5537
Level: single treatment within block Iteration: 16000 Score: 36.81988 Best: 36.81988 Since Improvement: 6537
Level: single treatment within block Iteration: 17000 Score: 36.81988 Best: 36.81988 Since Improvement: 7537
Level: single treatment within block Iteration: 18000 Score: 36.81988 Best: 36.81988 Since Improvement: 8537
Level: single treatment within block Iteration: 19000 Score: 36.81988 Best: 36.81988 Since Improvement: 9537
Early stopping at iteration 19463 for level single treatment within block
factorial_resultOptimised Experimental Design
----------------------------
Score: 36.81988
Iterations Run: 19464
Stopped Early: TRUE
Treatments: A1-B1, A1-B2, A1-B3, A2-B1, A2-B2, A2-B3, A3-B1, A3-B2, A3-B3, A4-B1, A4-B2, A4-B3, A5-B1, A5-B2, A5-B3, A6-B1, A6-B2, A6-B3, A7-B1, A7-B2, A7-B3, A8-B1, A8-B2, A8-B3
Seed: 112 Output of the Optimisation
The output summarises the optimisation of the factorial interaction treatment (e.g. "A1-B2") across the spatial layout. The reported score is for the whole design after optimisation, and the returned design_df contains the updated treatment allocation.
Because the treatment column encodes multiple factors, it can be helpful to split it back into its component factors (e.g. treatment_a and treatment_b) when inspecting the result. This lets you check whether the optimisation improved not only the interaction layout, but also the balance/adjacency patterns of the main effects.
str(factorial_result)List of 8
$ design_df :Classes 'design' and 'data.frame': 72 obs. of 8 variables:
..$ row : int [1:72] 1 1 1 2 2 2 3 3 3 4 ...
..$ col : int [1:72] 1 2 3 1 2 3 1 2 3 1 ...
..$ treatment : chr [1:72] "A6-B1" "A8-B2" "A5-B3" "A1-B2" ...
..$ row_block : num [1:72] 1 1 1 1 1 1 1 1 1 1 ...
..$ col_block : num [1:72] 1 1 1 1 1 1 1 1 1 1 ...
..$ block : num [1:72] 1 1 1 1 1 1 1 1 1 1 ...
..$ treatment_a: chr [1:72] "A1" "A1" "A1" "A2" ...
..$ treatment_b: chr [1:72] "B1" "B2" "B3" "B1" ...
..- attr(*, "out.attrs")=List of 2
.. ..$ dim : Named int [1:2] 24 3
.. .. ..- attr(*, "names")= chr [1:2] "row" "col"
.. ..$ dimnames:List of 2
.. .. ..$ row: chr [1:24] "row= 1" "row= 2" "row= 3" "row= 4" ...
.. .. ..$ col: chr [1:3] "col=1" "col=2" "col=3"
$ score : num 36.8
$ scores : num [1:19464] 131 141 139 141 153 ...
$ temperatures : num [1:19464] 100 99.9 99.8 99.7 99.6 ...
$ iterations_run: num 19464
$ stopped_early : logi TRUE
$ treatments : chr [1:24] "A1-B1" "A1-B2" "A1-B3" "A2-B1" ...
$ seed : num 112
- attr(*, "class")= chr [1:2] "design" "list"Visualise the Output
treatments <- strsplit(as.character(factorial_result$design_df$treatment), "-") |>
unlist() |>
matrix(ncol = 2, byrow = TRUE)
factorial_result$design_df[c("treatment_a", "treatment_b")] <- treatments
pa <- autoplot(factorial_result, treatments = "treatment_a")
pb <- autoplot(factorial_result, treatments = "treatment_b")
p <- autoplot(factorial_result)
p + pa + pb + plot_layout(ncol = 3)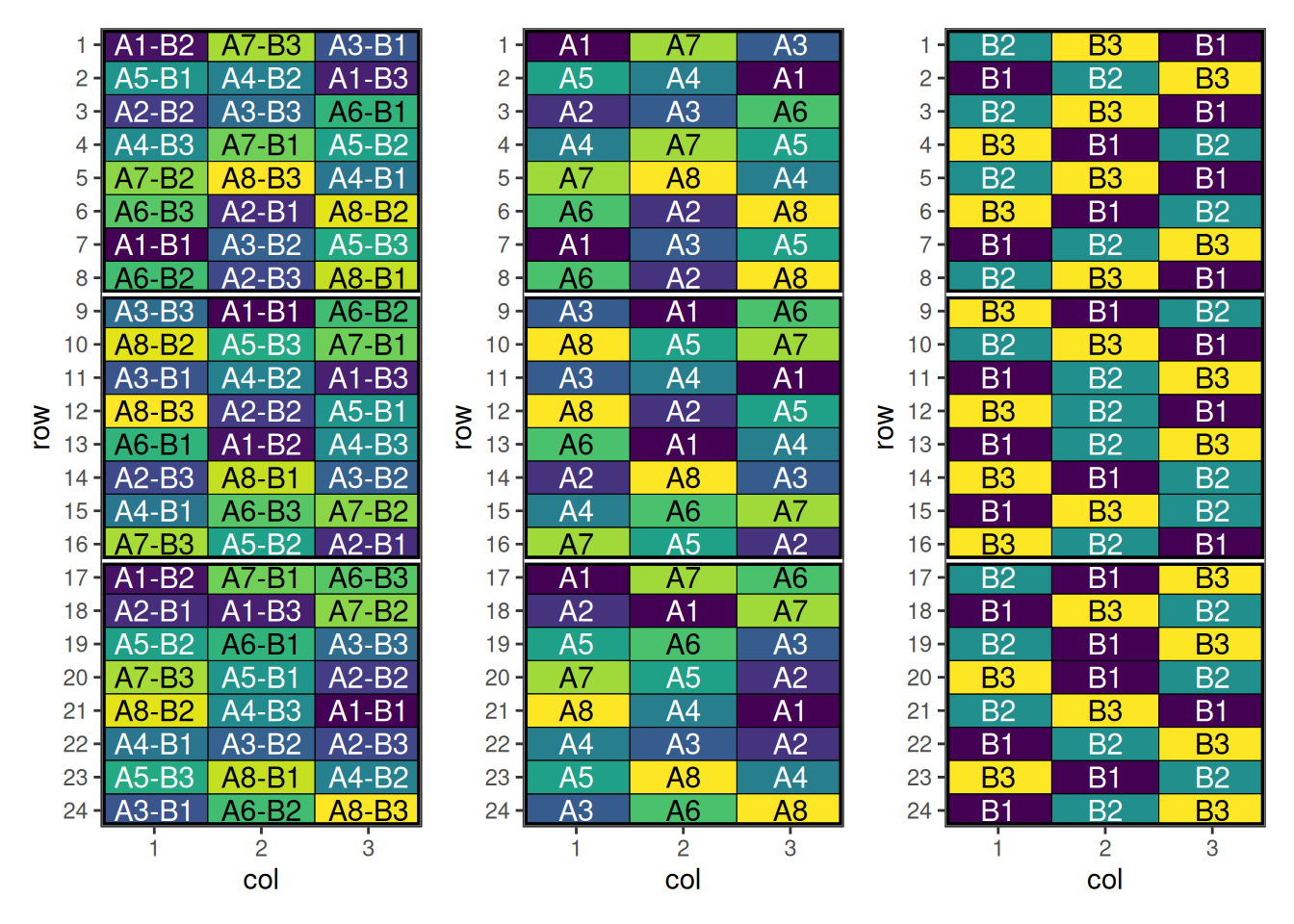
This design has now been optimised for both main and interaction effects.
Spatial Design Considerations
Field Shape and Orientation
Neighbour Effects
Using speed Effectively
- Set appropriate parameters: Balance optimisation time with improvement
- Visualise designs: Always plot layouts before implementation
- Compare alternatives: Test multiple blocking strategies
- Validate results: Check constraint satisfaction and efficiency factors
Conclusion
Further Reading
Related Vignettes
This vignette demonstrates the versatility of the speed package for agricultural experimental design. For more advanced applications and custom designs, consult the package documentation and additional vignettes.